harold fellermann
Bio-Nano-Technology
Optimizing nucleic acid sequences for a molecular data recorder
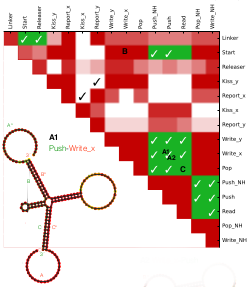
We recently reported the design for a DNA nano-device that can record and store molecular signals. Here we present an evolutionary algorithm tailored to optimising nucleic acid sequences that predictively fold into our desired target structures. In our approach, a DNA device is first specified abstractly: the topology of the individual strands and their desired foldings into multi-strand complexes are described at the domain-level. Initially, this design is decomposed into a set of pairwise strand interactions. Then, we optimize candidate domains, such that the resulting sequences fold with high accuracy into desired target structures both (a) individually and (b) jointly, but also (c) to show high affinity for binding desired partners and simultaneously low affinity to bind with any undesired partner. As optimization heuristic we use a genetic algorithm that employs a linear combination of the above scores. Our algorithm was able to generate DNA sequences that satisfy all given criteria. Even though we cannot establish the theoretically achievable optima (as this would require exhaustive search), our solutions score 90% of an upper bound that ignores conflicting objectives. We envision that this approach can be generalized towards a broad class of toehold-mediated strand displacement systems.
J. Kozyra, H. Fellermann, B. Shirt-Ediss, A. Lopiccolo & N. Krasnogor, Proceedings of the Genetic and Evolutionary Computation Conference GECCO, 2017
Designing uniquely addressable bio-orthogonal synthetic scaffolds for DNA and RNA origami
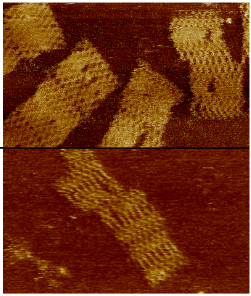
Nanotechnology and synthetic biology are rapidly converging, with DNA origami being one of the leading bridging technologies. DNA origami was shown to work well in a wide array of biotic environments. However, the large majority of extant DNA origami scaffolds utilize bacteriophages or plasmid sequences thus severely limiting its future applicability as bio-orthogonal nanotechnology platform. In this paper we present the design of biologically inert (i.e. "bio-orthogonal") origami scaffolds. The synthetic scaffolds have the additional advantage of being uniquely addressable (unlike biologically derived ones) and hence are better optimised for high-yield folding. We demonstrate our fully synthetic scaffold design with both DNA and RNA origamis and describe a protocol to produce these bio-orthogonal and uniquely addressable origami scaffolds.
J. Kozyra, A. Ceccarelli, E. Torelli, A. Lopiccolo, J. Gu, H. Fellermann, U. Stimming & N. Krasnogor, ACS Synthetic Biology, 2017
In vitro implementation of a stack data structure based on DNA strand displacement
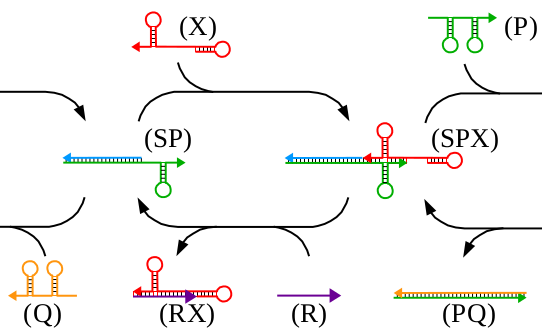
We present an implementation of an in vitro signal recorder based on DNA assembly and strand displacement. The signal recorder implements a stack data structure in which both data as well as operators are represented by single stranded DNA "bricks". The stack grows by adding push and write bricks and shrinks in last-in-first-out manner by adding pop and read bricks. We report the design of the signal recorder and its mode of operations and give experimental results from capillary electrophoresis as well as transmission electron microscopy that demonstrate the capability of the device to store and later release several successive signals. We conclude by discussing potential future improvements of our current results.
H. Fellermann, A. Lopiccolo, J. Kozyra & N. Krasnogor, Lect. Notes. Comp. Sci. 9726:87-98, 2016
Formalising Modularisation and Data Hiding for Synthetic Biology
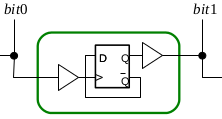
Biological systems employ compartmentalisation and other co-localisation strategies in order to orchestrate a multitude of biochemical processes by simultaneously enabling "data hiding" and modularisation. This paper presents recent research that embraces compartmentalisation and co-location as an organisational programmatic principle in synthetic biological and biomimetic systems. In these systems, artificial vesicles and synthetic minimal cells are envisioned as nanoscale reactors for programmable biochemical synthesis and as chassis for molecular information processing. We present P systems, brane calculi, and the recently developed chemtainer calculus as formal frameworks providing data hiding and modularisation and thus enabling the representation of highly complicated hierarchically organised compartmentalised reaction systems. We demonstrate how compartmentalisation can greatly reduce the complexity required to implement computational functionality, and how addressable compartments permits the scaling-up of programmable chemical synthesis.
H. Fellermann, M. Hadorn, R. Füchslin & N. Krasnogor, ACM J. Emerg. Tech. Comp. Sys 11(3):24, 2014
H. Fellermann & N. Krasnogor, Lect. Notes Comp. Sci. 8493:173-182, 2014
Programming Chemistry in DNA Adressable Bioreactors
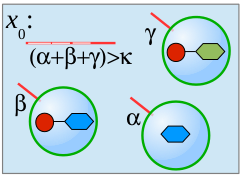
We present a formal calculus, termed chemtainer calculus, able to capture the complexity of compartmentalized reaction systems such as populations of possibly nested vesicular compartments. Compartments contain molecular cargo as well as surface markers in the form of DNA single strands. These markers serve as compartment addresses and allow for their targeted transport and fusion, thereby enabling reactions of previously separated chemicals. The overall system organization allows for the setup of programmable chemistry in microfluidic or other automated environments. We introduce a simple sequential programming language whose instructions are motivated by state-of-the-art microfluidic technology. Our approach integrates electronic control, chemical computing, and material production in a unified formal framework that is able to mimick the integrated computational and constructive capabilities of the subcellular matrix. We provide a non-deterministic semantics of our programming language that enables us to analytically derive the computational and constructive power of our machinary. This semantics is used to derive the sets of all constructable chemicals and supermolecular structures that emerge from different underlying instruction sets. Since our proofs are constructive, they can be utilized to automatically infer control programs for the construction of target structures from a limited set of resource molecules. Finally, we present an example of our framework from the area of oligosaccharide synthesis.
H. Fellermann & L. Cardelli, R. Soc. Interface 11(99), 2014
The MATCHIT Automaton: Exploiting Compartmentalization for the Synthesis of Branched Polymers
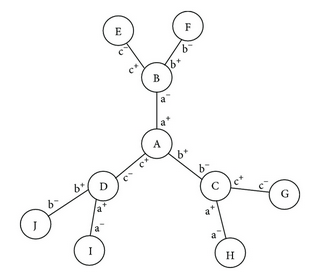
We propose an automaton, a theoretical framework that demonstrates how to improve the yield of the synthesis of branched chemical polymer reactions. This is achieved by separating substeps of the path of synthesis into compartments. We use chemical containers (chemtainers) to carry the substances through a sequence of fixed successive compartments. We describe the automaton in mathematical terms and show how it can be configured automatically in order to synthesize a given branched polymer target. The algorithm we present finds an optimal path of synthesis in linear time. We discuss how the automaton models compartmentalized structures found in cells, such as the endoplasmic reticulum and the Golgi apparatus, and we show how this compartmentalization can be exploited for the synthesis of branched polymers such as oligosaccharides. Lastly, we show examples of artificial branched polymers and discuss how the automaton can be configured to synthesize them with maximal yield.
M. Weyland, H. Fellermann, M. Hadorn, D. Sorek, D. Lancet, S. Rasmussen, & R. M. Füchslin, Comput. Math. Met. Med. 467428, 2013
Specific and Reversible DNA-directed Self-Assembly of Oil-in-Water Emulsion Droplets
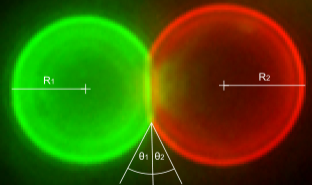

Higher-order structures that originate from the specific and reversible DNA-directed self-assembly of microscopic building blocks hold great promise for future technologies. Here, we functionalized biotinylated soft colloid oil-in-water emulsion droplets with biotinylated single-stranded DNA oligonucleotides using streptavidin as an intermediary linker. We show the components of this modular linking system to be stable and to induce sequence-specific aggregation of binary mixtures of emulsion droplets. Three length scales were thereby involved: nanoscale DNA base pairing linking microscopic building blocks resulted in macroscopic aggregates visible to the naked eye. The aggregation process was reversible by changing the temperature and electrolyte concentration and by the addition of competing oligonucleotides. The system was reset and reused by subsequent refunctionalization of the emulsion droplets. DNA-directed self-assembly of oil-in-water emulsion droplets, therefore, offers a solid basis for programmable and recyclable soft materials that undergo structural rearrangements on demand and that range in application from information technology to medicine.
M. Hadorn, E. Bönzli, K. T. Sørensen, H. Fellermann, P. Eggenberger-Hotz & M. Hanczyc, Proc. Nat. Acad. Sci. USA 109(47), 2012
DNA Self-Assembly and Computation Studied with a Coarse-grained Dynamic Bonded Model
We study DNA self-assembly and DNA computation using a coarse-grained DNA model within the directional dynamic bonding framework [C. Svaneborg, Comp. Phys. Comm. 183, 1793 (2012)]. In our model, a single nucleotide or domain is represented by a single interaction site. Complementary sites can reversibly hybridize and dehybridize during a simulation. This bond dynamics induces a dynamics of the angular and dihedral bonds, that model the collective effects of chemical structure on the hybridization dynamics. We use the DNA model to perform simulations of the self-assembly kinetics of DNA tetrahedra, an icosahedron, as well as strand displacement operations used in DNA computation.
C. Svaneborg, H. Fellermann & S. Rasmussen, Lect. Notes in Comput. Sc. 7433, 123, 2012
